GPCRdb
G protein-coupled receptors (GPCRs) are cell-surface receptors mediating the responses of 2/3rds of human hormones1 and 1/3rd of
drugs2.
Each GPCR can bind several transducers, G proteins, GPCR kinases (GRKs) and arrestins
leading to distinct intracellular signaling networks and functional outcomes.
Data
- 805 Human proteins
- 42,021 Species orthologs
- 3,019 Generic residues
- 1,716 GPCR structures
- 880 Refined structures
- 1,601 GPCR structure models
- 634 Physiological ligand-GPCR structure models
- 546 Drugs
- 173 Compounds in trial
- 121 Drug targets
- 683 Disease indications
- 222,036 Ligands
- 527 Physiological ligands
- 499,650 Ligand bioactivities
- 35,588 Ligand site mutations
- 66,281 Ligand interactions
Latest release
March 20, 2025- Complete redesign of the Drug section
- New improved and curated Drug data (Nature Reviews Drug Discovery, 2025)
- Dedicated sections for agents/drugs, indications and drug targets
- Overview of the drugged GPCRome
- New structures
- Minor bug and performance fixes
All releases
Publications
GPCRdb in 2025: adding odorant receptors, data mapper, structure similarity search and models of physiological ligand complexes.
Nucleic Acids Research, 2024
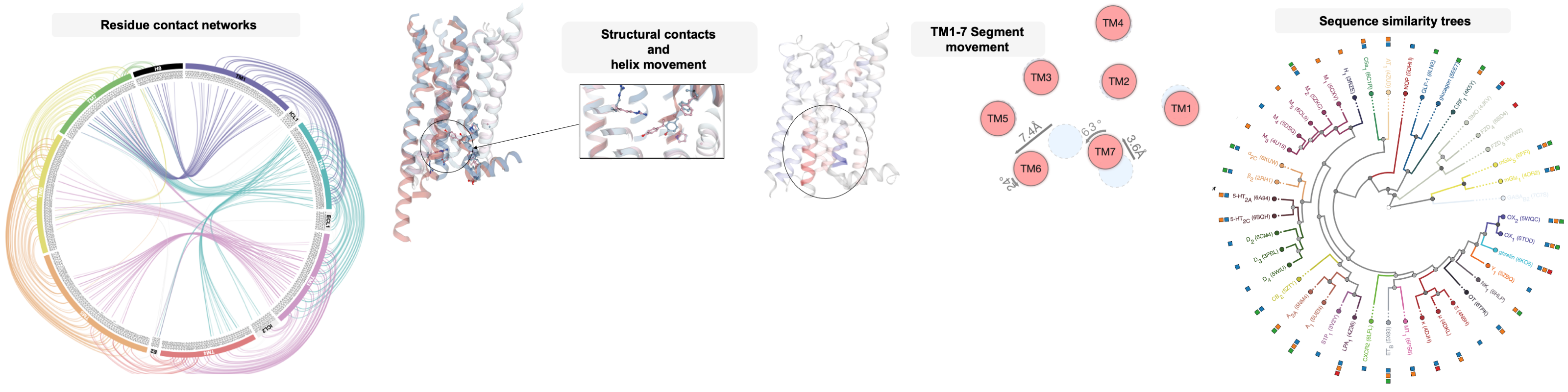
An online GPCR structure analysis platform.
Nature Structural & Molecular Biology, 2021
GPCR drug discovery: new agents, targets and indications.
Nature Reviews Drug Discovery, 2025
Generic GPCR residue numbers - aligning topology maps while minding the gaps.
Trends in pharmacological sciences, 2015